Imaging the invisible: how can research software and imaging techniques help scientists study the things we can’t see?
From microscopic plankton to individual atoms, the subjects of many scientific studies need special devices to be seen. Scientific imaging devices, like electron microscopes, have become increasingly powerful in recent years, allowing scientists to study their subjects in even more detail. These devices need to be partnered with software to allow the images they produce to be analysed. Dr Joanna Leng, from the University of Leeds in the UK, is a research software engineer who designs and develops the software that allows scientific imaging devices to be used to their full potential.
TALK LIKE A RESEARCH SOFTWARE ENGINEER
Scientific imaging — the process of capturing or creating images of scientific phenomena. This often involves the use specialist devices such as microscopes
Scientific model — a conceptual or mathematical representation of a real-world phenomenon that allows scientists to study the phenomenon in more detail
Scientific simulations — an advanced type of computational model that not only represents a real-world phenomenon, but aims to predict how the phenomenon might change under different conditions or parameters
Software — a set of instructions, scripts or programmes that are used to operate computers and perform specific tasks
Synchrotron — a type of particle accelerator
From miniscule sub-atomic particles to gargantuan black holes, the world of science deals with a dramatic range of sizes. Atoms are millions of times smaller than a grain of sand, and the visible universe stretches out for 13.5 billion lightyears. This poses the question: How do you study things that you cannot see?
This quandary has led to the creation of many imaging devices that allow scientists to work with such extreme scales. For example, using electron microscopes, we can now capture images at an atomic resolution in which each pixel is less than the size of an atom. At the other end of the scale, NASA’s recently-launched James Webb Space Telescope will be able to see to the very edge of the visible Universe, showing us the stars and galaxies that formed shortly after the Big Bang.
As scientists gaze deeper and further into the extreme reaches of our Universe, the need for more powerful and sophisticated imaging devices grows. But these devices do not work by themselves. They need to be operated by experts who know how to use them and paired with software that allows scientists to analyse the images they produce. Dr Joanna Leng, from the University of Leeds in the UK, is a research software engineer who specialises in developing software for new imaging techniques.
Why is scientific imaging so important?
“Scientific images and videos allow us to see things that we cannot see with our unaided-eyes,” explains Joanna. Objects such as cells and crystals, and the processes through which they are created, exist at microscopic scales, and scientific imaging allows scientists to study them. With the help of scientific imaging, researchers can conduct experiments to test their hypotheses and build new theories about how the Universe works. “These are the first two pillars of scientific discovery,” says Joanna. “Theory and experimentation.”
Joanna believes the third pillar of scientific discovery is computer simulation, a technique which is becoming increasingly popular. Scientists can now turn their theories into mathematical models, which can then be expressed in software as simulations. After running the simulations, scientists can study their outputs and compare them to experimental results. For example, the outputs can be visualised using computer graphics and compared directly to the images captured in experiments. This can lead to new discoveries or the development of new theories that can advance our understanding of the Universe.
Over the last couple of decades, a new technique has been developed that merges computer simulation with scientific imaging techniques. Called image-based modelling, the technique turns images into 2D or 3D models, which can be a complex process. Simulations and images handle data in individual units, whereas in reality, space is continuous. “The process of turning continuous space or real objects into a set of discrete units is called mesh generation,” explains Joanna. Since computer simulations were first introduced, computers have become much more powerful, allowing them to create meshes which are larger, more detailed and more realistic. “This is an exciting change as it has significantly increased the speed and accuracy of scientific discovery,” says Joanna.
What is research computing?
Reference
https://doi.org/10.33424/FUTURUM314
Photo Lawrence Molloy
As imaging systems and modelling techniques have become more powerful and complex, they have become harder to operate and the results they produce harder to interpret. As a result, a new discipline, known as research computing, has emerged to apply computers, not just software, to research including to help scientists capture images, construct models, which are turned into simulations, and analyse results.
Research computing is a sub-discipline of computer science. However, it is needed across a wide range of academic fields. As scientific imaging and simulation has become more widespread, the need for research computing has created synergies between disciplines that, traditionally, had little in common. The techniques used to analyse images and the software used to create them are often very similar, no matter which discipline a scientist is working in.
Many national research facilities have sophisticated imaging systems that often require specific expertise to operate. For example, Diamond Light Source is home to the UK’s national synchotron. When researchers visit the facility to capture images, they are supported by scientists, including research software engineers who help them process their data and access software.
Although research computing support is present at most national research facilities, other organisations in the UK may not offer similar support. For example, when researchers return from a national research facility to their home university or business, they may lack the equipment or expertise needed to progress with their work. The lack of research computing at these organisations means that it can take years for researchers to analyse their data in full.
How can the UK improve its research computing?
“There is a need for expertise in the UK that allows techniques and knowledge to be shared and transferred across all academic disciplines,” explains Joanna. There is an opportunity for budding computer scientists to make themselves invaluable to the UK’s scientific community.
Research computing is slowly becoming recognised as a distinct profession, with its own set of practices, but Joanna believes a lack of funding is limiting its progress. “Funding needs to be sustained over a long period so that people can develop the skills, jobs and then careers in research computing,” she says. Without the development of research computing expertise, the UK’s scientific community risks falling behind. One area of research computing that is in high demand is research software engineering.
Why do we need research software engineers?
“There is little we can do in the modern world without computers,” says Joanna. “This is true for research, too.” Research software engineers design and adapt the software that scientists need when working with complex scientific imaging and simulation tools. This software is needed to collect and analyse the data from images, as well as to create simulations and compare their results.
Because computational methods originated in the natural sciences, some disciplines, such as chemistry and physics, have lots of research software at their disposal. Other disciplines, like the social sciences, have much less, although they are starting to catch up in areas which are important to social media and its regulation, like computational linguistics and social network analysis.
Nowadays, all areas of research have a need for software, and this need will only increase in the future. As such, research software engineering has the potential to become one of the most important scientific disciplines in the coming years. As new computational technologies are developed, imaging systems and simulation techniques are likely to become even more complex than they are now. These advancements in technology will need to be matched by advancements in software, resulting in an even more pressing need for research software engineers.
 DR JOANNA LENG
DR JOANNA LENG
School of Computing, University of Leeds, UK
Field of research: Research Software Engineering
Research project: Developing new software for new imaging techniques
Funder: Engineering and Physical Sciences Research Council (EPSRC)
This work was supported by the EPSRC grant number EP/R025819/1 (gow.epsrc.ukri.org/NGBOViewGrant.aspx?GrantRef=EP/R025819/1)
Joanna’s research
Joanna is a senior research software engineer (RSE) at the University of Leeds, holding a prestigious RSE Fellowship from the Engineering and Physical Sciences Research Council (EPSRC). It is her responsibility to support other researchers at the university when their projects require complex imaging devices and modelling software. As an RSE, her work spans many different disciplines, so she gets to work on lots of exciting projects.
Chemistry with Dr Nicole Hondow and Stuart Micklethwaite
Joanna is working with Nicole and Stuart on a project that involves an analytic technique called Energy-Dispersive X-ray spectroscopy (EDX). This technique is used to analyse the elements that make up a sample. To do this, they use an electron microscope to take a very detailed image of the sample.
The electron microscope fires an electron beam which is scanned across the surface of the sample. When the atoms within the sample are excited by the electron beam, they emit electrons, which is recorded as an image, and X-rays. As each atom emits unique X-rays, Joanna is able to determine the elemental composition of the sample. Once the surface layer of the sample has been mapped, it is removed, allowing the layer underneath to be mapped in the same way. Slowly, a 3D map of the sample is built up. “As you can imagine, the size of these images is huge, and they require large computational resources to process and analyse,” says Joanna.
This technique enables Joanna to study samples from the millimetre to the atomic scale, meaning that she can consider the chemical states of materials in much finer detail. By using the EDX technique, Joanna, Nicole and Stuart can study the properties of well-known materials and apply their findings to creating new ones.
Chemical engineering with Professor Sven Schroeder
Joanna is working with Sven, an expert in chemical and process engineering, on a technique known as continuous tubular crystallisation (CTC). This is a new technique that has enormous potential to improve efficiency and consistency in industrial processes and reduce costs.
Sven and one of his PhD students have been working at Diamond Light Source to develop new ways of producing video footage of CTC using X-Rays. When first experimenting with video footage, the features were so faint that they could hardly make out any crystals, at all. Fortunately, Joanna was able to help them use a new video processing technique called Eulerian Video Magnification (EVM). This enabled Sven and his team to study the crystals in much more detail and track their growth. In the new footage, they were able to identify a process a called ‘oiling out’ which is bad for chemical processes and indicates impurities in the crystals. Using more sophisticated imaging systems to identify problems like this can help to improve industrial processes like CTC.
Joanna and Jonathan Pickering, an RSE employed by the EPSRC fellowship, developed software that allows EVM video footage to be analysed and annotated manually. She is also helping Sven to supervise a new PhD student who is using this software to improve the CTC imaging technique so that the EVM post-processing technique is no longer necessary. Once this is done, the CTC video footage will be accessible to artificial intelligence image analysis techniques.
Biology with Professor Michelle Peckham and Dr Alistair Curd
Michelle and Alistair develop and use new imaging techniques. Joanna has worked closely with them to help develop and improve the software that these imaging techniques use. This requires a lot of work, including updating the code, running tests and producing reports. As a result, Michelle and Alistair’s software is now easier to install and use.
ABOUT RESEARCH SOFTWARE ENGINEERING
Research software engineering (RSE) is a relatively new discipline that involves developing and using the software that is needed for scientific research. Software is needed in almost all scientific fields, so research software engineers (RSEs) are in high demand.
What opportunities will be open to the next generation of RSEs?
As more disciplines embrace computational and imaging techniques, funders and senior university administrators will recognise the growing need for RSEs. As a result, funding will increase, and organisational structures will change to reflect the needs of academic research. This will increase the opportunities for RSEs and allow them to develop new and innovative software for more research areas. They will also have access to better infrastructure and laboratory equipment, such as supercomputers and quantum computers.
While software will be designed by RSEs for research, it may have unexpected applications for wider society. “Don’t forget, the World Wide Web started as a way for scientists at CERN to share information, and Google was devised by two PhD students while they were at Stanford University,” says Joanna. As RSE accrues better recognition as a profession, with a clear career path, the opportunities will continue to expand.
Joanna’s top tips
1. The best way to find out about the opportunities in your area is to talk to a research software engineer! Attend national research facility open days and events, and talk to as many scientists as you can.
2. If you can’t attend open days, sign-up for universities’ and colleges’ social media updates and newsletters.
Pathway from school to RSE
• RSEs use computers and software to solve problems, so you will need to be curious and enjoy tackling challenges.
• You will also need to develop your people skills so that you can communicate well. It is important that you fully understand the problems that researchers are trying to solve.
• Most RSEs have a degree and/or PhD in their chosen area, such as maths, physics or chemistry. Whilst taking part in research projects, they will have had to develop software and learn about different software engineering techniques. These skills help them to transition into RSEs.
• Modelling and simulations are often required for complex problems, such as climate change, and are employed by government agencies. RSEs who develop this software will likely have qualifications in maths, statistics, physics and/or computer science.
• It is important to study maths to at least A-level/post 16-years-old. Learning computing and computer programming will also be beneficial.
Scientific imaging in physics
Sarah’s research

Dr Sarah Harris
Theoretical physicist,
School of Physics and Astronomy,
University of Leeds
Sarah’s interests are in understanding biology. “I want to understand how proteins and DNA use the laws of physics to perform their functions,” says Sarah. “To do that, I use computer simulations of these proteins and DNA.”
Sarah models individual proteins or small pieces of DNA. These structures are so tiny that they often change shape due to fluctuations in thermal energy. The models and simulations that Sarah and her team create allow them to study these changes in atomic detail.
One of the biggest applications of Sarah’s models is in drug design. Drugs bind to proteins using a lock and key mechanism, in which the drug is the key, and the protein molecule is the lock. “You need to have the key that’s the right shape to go into the lock for the interaction to proceed,” explains Sarah. “The difficulty with proteins is that they’re always changing shape, and you need to try lots of different keys against lots of different shapes of this lock to see which ones fit most of the time.” The models that Sarah creates help to solve this problem.
Computational tools are used throughout the pharmaceutical industry. Despite this, they still suffer from limitations. “Even with the fastest and biggest supercomputers and the best software, we still can’t run simulations that are long enough to predict how strongly a drug will bind to a protein,” explains Sarah. Currently, lots of research is focused on developing faster and more powerful software.
Sarah’s top tips
1. Be curious and open-minded.
2. Don’t give up when things go wrong.
3. Don’t expect everything to work the first time you try it.
Scientific imaging in in cell biology
Michelle’s Research

Professor Michelle Peckham
Faculty of Biological Sciences,
University of Leeds
Michelle’s research is focused on the cytoskeleton, a network of interlinking protein filaments found in cells. The cytoskeleton helps cells maintain their shape and move around. It also facilitates the internal transport of organelles and other structures present within the cell. For example, a motor protein called myosin uses the cytoskeleton as a track to navigate around the cell. Without the cytoskeleton, it would not be able to perform its role as a key component of muscle contraction. Michelle’s research ranges from basic research into how myosins perform their functions to more complex studies of how mutations in cytoskeletal proteins cause disease.
How does Michelle use scientific imaging in her research?
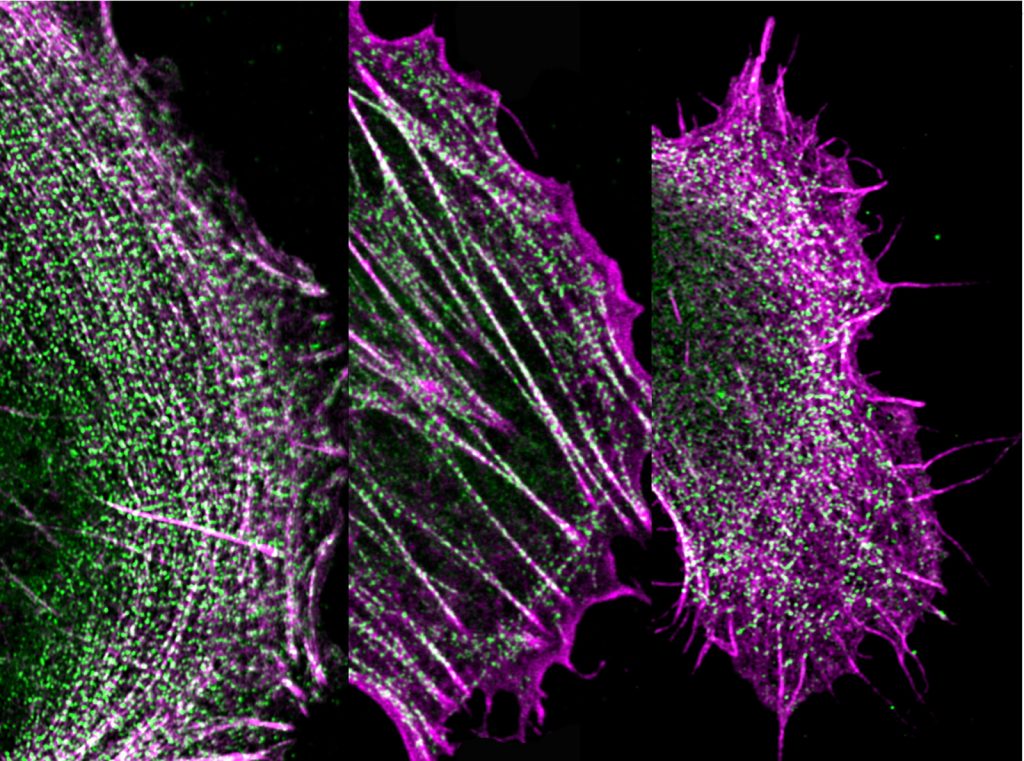
Michelle uses a variety of complex imaging techniques when conducting her research. For example, cryo-electron microscopy, which she uses to study the structure of myosins, involves freezing samples to below -150 °C and imaging them using beams of electrons. Another imaging technique, called super-resolution light microscopy, has allowed Michelle to produce detailed images of the cytoskeleton within different kinds of cell. These images have enabled her to investigate a range of biological questions including how viruses move around in cells and how the cytoskeleton is organised within specialised cells such as muscle cells.
Michelle has collaborated with various other scientists to develop a type of microscopy known as dSTORM. This system allows her to see how proteins are organised within tiny structures that would not be visible with standard fluorescent microscopes. For example, some proteins found within specialised structures in muscle cells are just 20nm apart. Objects of this size are well below the capabilities of standard fluorescent microscopes.
Why is dSTORM microscopy better than fluorescent microscopy?
Fluorescent microscopy uses a high-intensity source of light to excite fluorescent molecules within a sample. When these molecules are excited, they emit a lower energy light with a longer wavelength, which generates the final image. The problem with this technique is that all the molecules emit their fluorescence at the same time. This makes it impossible to identify molecules that are too close together, as their fluorescence overlaps and creates a blurry image.
In the dSTORM system, a special buffer is used that encourages most of the fluorescent molecules to enter a non-fluorescent state. This means that at any one time, only a small subset of the molecules are fluorescent and these are far enough apart to be imaged as single molecules, and their locations identified precisely. After a short period of time, this subset of fluorescent molecules bleach and a new subset of molecules becomes fluorescent, imaged and identified. This ‘blinking behaviour’ continues over a long period of time and generates a large number of localisations, which are combined to generate a much more accurate and detailed image.
What is the dSTORM system used for?
dSTORM is used in laboratories all over the world to determine how proteins are organised in many different cellular structures. For example, the dSTORM system has been used to analyse the organisation of proteins within the nuclear pore, a channel which regulates the movement of molecules between the nucleus and the rest of the cell. Michelle has also used dSTORM to produce images of a protein that causes diseases relating to cilia, which are small antennae found on the cell surface. Using this technique, she was able to demonstrate, for the first time, how this protein is organised.
Does the dSTORM system have real-world applications?
Producing images of proteins that cause diseases may point the way to novel treatments and cures. For example, understanding how proteins are organised in muscle cells is important for understanding heart and skeletal muscle disease. Many of these proteins can undergo mutations that are caused by disease. Investigating how they are organised could be key to understanding the molecular basis of such diseases.
Alistair’s research

Dr Alistair Curd
Faculty of Biological Sciences,
University of Leeds
Alistair conducts research that seeks to improve optical microscopy techniques and use them to investigate biological systems and diseases. Alistair develops super-resolution microscopes and new data analysis techniques which he uses to collaborate on experiments with other biologists and medics.
Recent advances in microscopy have enabled microscopes to see down to extraordinarily tiny scales. Unfortunately, this has resulted in the systems becoming much slower and more complicated. Because of their complexity, some of these highly-powerful microscopes can still miss molecules when producing an image. Alistair is developing software for super-resolution microscopes that can identify the patterns that dictate how molecules are arranged in cells. This software would then be able to use these patterns to work out the positions of the molecules missing from an image.
Alistair’s software examines pairs of molecules that have been captured in images by a super-resolution microscope. It then calculates the relative position of each pair by observing the distance and direction between them. The software finds the relative positions for many molecule pairs and is then able to identify patterns within a sample. The software can then compare the pattern of relative positions with the positions that would be expected in specific circumstances. From this comparison, the software can identify the hypothesis which best matches the data.
He is currently working with a cancer surgeon to investigate bowel cancer. They are trying to discover how bowel cancer develops and why some treatments are more effective than others. Alistair’s software will enable them to study the important molecules in greater detail than would otherwise have been possible.
ABOUT CELL BIOLOGY
Cell biology is a branch of biology that studies the structure, function and behaviour of cells. Due to the microscopic nature of the subjects, cell biologists constantly make use of imaging systems.
What is rewarding about cell biology?
“Cell biology is an area of research full of unsolved problems,” says Michelle. “As a problem solver, for me, gaining an understanding of how things work for the first time is extremely rewarding.” Michelle also enjoys developing the research methods which enable her to conduct new types of experiments and discover new things.
How will scientific imaging in cell biology develop?
An as yet undeveloped technique called correlative light and electron microscopy is the next big challenge. It combines light microscopy with electron microscopy to overcome some of the limitations of cryo-electron microscopy. CryoEM allows researchers to study the structure of proteins when they are isolated. However, it is much harder to study proteins in their natural environment within a cell as specific proteins cannot be labelled. On the other hand, fluorescent microscopy is able to label and image proteins within cells.
“Combining the two approaches should help us to see protein structure and organisation within cells to extraordinarily high resolutions,” says Michelle. She hopes that this new technique will pave the way for new discoveries into how proteins interact with each other, and how these interactions are disrupted due to diseases.
Pathway from school to cell biology
• Study as many science subjects as you can at school. Michelle recommends studying physics, chemistry, maths and biology.
• Science is becoming much more interdisciplinary, so the more disciplines you learn about, the better!
• Michelle says, “Find something that you enjoy and be persistent with it. Not everything is going to go right first-time round.”
Scientific imaging in chemistry
Nicole’s Research

Dr Nicole Hondow
School of Chemical and Process Engineering,
University of Leeds
Nicole’s research focuses on material characterisation, which is the process of probing and measuring the properties of different materials. These properties might include the chemical make-up of a material or its physical and mechanical structures.
Nicole’s research focuses primarily on using electron microscopes to study nanomaterials.
Nicole uses electron microscopes to study what her samples are made of and what shapes and sizes they are, as these properties influence their potential uses. Engineered nanomaterials are being utilised in a number of different areas including medicine, energy and the environment. For example, nanomaterials can be used to remove toxins and harmful substances, such as those found in car exhaust fumes, from the air.
What imaging techniques does Nicole use?
Nicole uses a Focused Ion Beam Scanning Electron Microscope (FIB-SEM) to image her samples. These microscopes use electrons (to image the sample) and ions (to alter the sample). The ion beam is used to remove layers of the sample so that the interior structure can be viewed and imaged. Nicole uses this technique to take successive images of a sample, probing deeper into its structure each time. She uses these images to build videos and 3D models of the samples, which she can then analyse.
What are the applications of Nicole’s research?
The FIB-SEM technique allows Nicole to study her samples in much more detail than more conventional microscopes would. “Being able to undertake nanoscale analysis inside materials helps us understand their structure,” says Nicole. “From this we can then look at what might make them better – how we can make the materials perform better or how to improve their design.”
Nicole’s aim is to discover new ways in which the design of her nanomaterial samples can be improved to maximise their performance in a range of different applications. Nicole hopes to expand her range of samples to include nanomaterials being used in the development of new medicines, foods and technologies that tackle environmental problems.
What opportunities are there for research software engineers in chemistry?
There are lots of opportunities to get involved in the development of new materials and chemical products, particularly those that benefit the environment and human health. As global issues such as climate change and the COVID-19 pandemic become increasingly prevalent, the need for innovative solutions will only increase.
Research software engineers will be a vital part of this process. Chemists like Nicole can use microscopes to generate lots of data and images; but they then need to be able to process and analyse them. Nicole relies on research software to help her truly understand the materials that she studies. As imaging techniques become more complex and powerful, the research software that accompanies them will need to improve at the same pace.
“It is always exciting to solve a problem or figure out any small part of some larger question,” says Nicole. Research software engineers are often at the heart of this problem-solving process.
Pathway from school to chemistry
• In addition to chemistry, maths is very important, but also consider physics, biology and computing as these can provide
great opportunities.
• Salters’ Institute provides lots of resources and organises events to help students explore chemistry and related careers. Its student page is a good starting point:
www.saltersinstitute.org/iam/i-am-a-secondary-school-student
• Nicole says, “Finding an area that you are passionate about is crucial. Find an area that you are truly interested in, and pursue it.”
Meet Lawrence

Lawrence Molloy
Artist in Residence, The Museum of the History of Science, Technology and Medicine at the University of Leeds
Lawrence is a sculptor, performer and curator who explores scientific concepts through creativity and play. He has worked with both Joanna and Nicole to create art based on their research projects. Working collaboratively and across disciplines allows Lawrence to explore the mechanisms of creativity and play in unexpected areas.
Lawrence has worked with museums, educators, scientists and historians to create a variety of artworks. “In each case, I have responded to the subject matter or technology,” says Lawrence. “But the hook has always been the excitement that my collaborators have for their field.” Some subjects may appear to be mundane or confusing at first glance, but when seen through the eyes of a specialist, they can become something magical. Often, these specialists have dedicated their lives to understanding and exploring something very specific, so their passion really shines through.
As a collaborative artist, Lawrence’s first job is to learn about his collaborator’s field. He needs to understand their subject area so that he can ask them meaningful questions that could spark creative and artistic ideas. Lawrence’s aim is to create art that allows audiences to experience scientific phenomena in a playful and engaging way. For example, he created a sculpture called Unit Cell to explore the mapping of crystal lattices. Unit Cell was made up of 125 beach balls arranged in a cubic grid, with each ball representing an atom. Lawrence and his collaborators, Professor Ben Whitaker, Dr Mike Nix and Dominic Hopkinson, fired sound waves at the balls to demonstrate how diffraction through a crystal lattice affects mapping techniques.
Lawrence met Nicole and her team at the Leeds Electron Microscopy and Spectroscopy Centre (LEMAS). It was important for Lawrence to experience the scanning process first-hand for three reasons. He explains, “Firstly, I started to see how exploring scales this small could feel like a journey through, and into, something alien. Secondly, I saw the skill it took to use, navigate and interpret results from these complex microscopes, and thirdly, I began to understand the limits and advantages of different microscopy techniques.”
Lawrence and Nicole had many discussions about how to visualise her data and the different strategies that they could use to communicate her complex scientific ideas. “Even though I knew the scales and had it repeatedly explained to me, I could not get my head around navigating or conceptualising at such small sizes,” says Lawrence. Another problem that Lawrence encountered was that the images produced by electron microscopes are already beautiful and captivating. He explains, “The question for me was, what do I bring to the party?””
To explore this question, Lawrence invited Nicole and her team over to his house for a curry. They discussed how Lawrence could facilitate the team’s ideas and Lawrence encouraged them to be wildly creative. He then returned to LEMAS and scanned images of the spices that they had used while cooking the curry. “The images that came out of this were interesting,” says Lawrence. “Some were even beautiful, but the subject matter, while fun, was not important.”
Lawrence is still exploring the question of how to turn data sets from microscopy into interactive sculptures. The process of creating these artworks can be long and challenging, but taking the time to truly understand his subject is a key part of Lawrence’s work. It is this involved approach that makes Lawrence’s sculptures so captivating as he is able to combine his collaborator’s knowledge and passion with his unique creativity.
Do you have a question for Joanna, Sarah, Michelle, Alistair, Nicole or Lawrence?
Write it in the comments box below and Joanna, Sarah, Michelle, Alistair, Nicole or Lawrence will get back to you. (Remember, researchers are very busy people, so you may have to wait a few days.)